
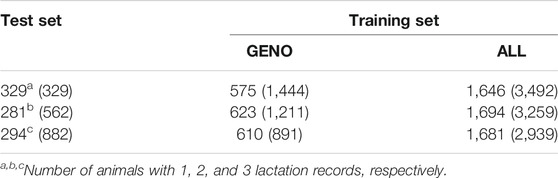
A total of 2,149 individuals from NAM populations were sequenced by exome capture and two sets of 15 SNP matrices (one for each family) were generated. Alignment of these sequences with the reference genome, AP13 (v3.0), revealed that up to 99% of the genomic sequences mapped to the reference genome. Genomic shotgun sequencing of 15 switchgrass NAM founder parental genomes at JGI produced 28-66 Gb high-quality sequence data. Dried biomass samples were also analyzed using prediction equations of NIRS at the Noble Foundation and for lignin content, S/G ratio, and sugar release characteristics at the NREL. Phenotypic data on plant height, tillering ability, regrowth, flowering time, and biomass yield were collected. All the progenies, founder parents, F1 parents (n=2350) were evaluated in replicated field trials at Ardmore, OK and Knoxville, TN.

The switchgrass NAM population consists of a total of 2000 genotypes from 15 families. Progenies form each family were randomly selected to develop the NAM population.
Ten genotypes from each of the 15 F1 families were then chain crossed. To develop a NAM population of switchgrass, 15 highly diverse genotypes with specific characteristics were selected from a diversity panel and crossed to a recurrent parent, AP13, a more » genotype selected for whole genome sequencing and parent of a mapping population.
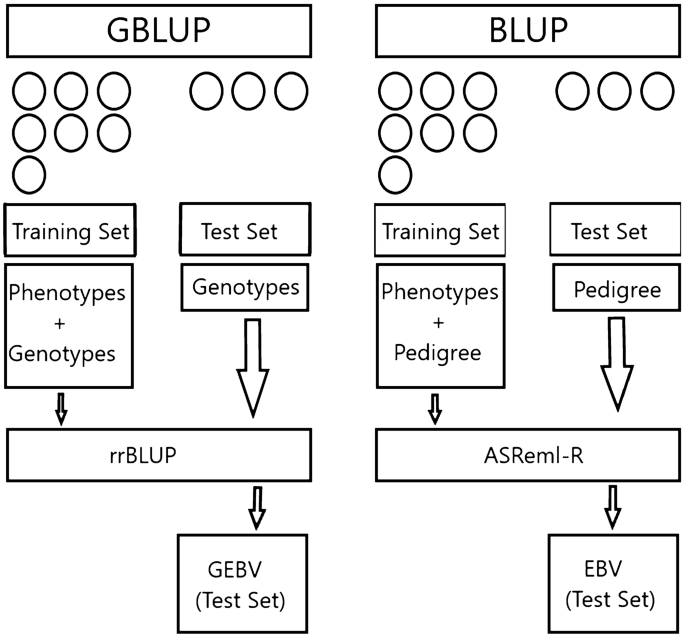
The nested association mapping (NAM) analysis combines the best features of both bi-parental and association analyses and can provide high power and high resolution in QTL detection and will ensure significant improvements in biomass yield and quality. Understanding the genetic basis of quantitative traits is essential to facilitate genome-enabled breeding programs. Switchgrass (Panicum virgatum L.) is a C4 grass with high biomass yield potential and a model species for bioenergy feedstock development. of California, Oakland, CA (United States) Sponsoring Org.: USDOE Office of Science (SC) OSTI Identifier: 1258157 Grant/Contract Number: AC02-05CH11231 Resource Type: Journal Article: Accepted Manuscript Journal Name: G3 Additional Journal Information: Journal Volume: 6 Journal Issue: 4 Journal ID: ISSN 2160-1836 Publisher: Genetics Society of America Country of Publication: United States Language: English Subject: 59 BASIC BIOLOGICAL SCIENCES genomic selection linkage disequilibrium exome capture bioenergy Panicum virgatum L GenPred shared data = , Research Service United States Department of Agriculture, Madison, WI, (United States)
